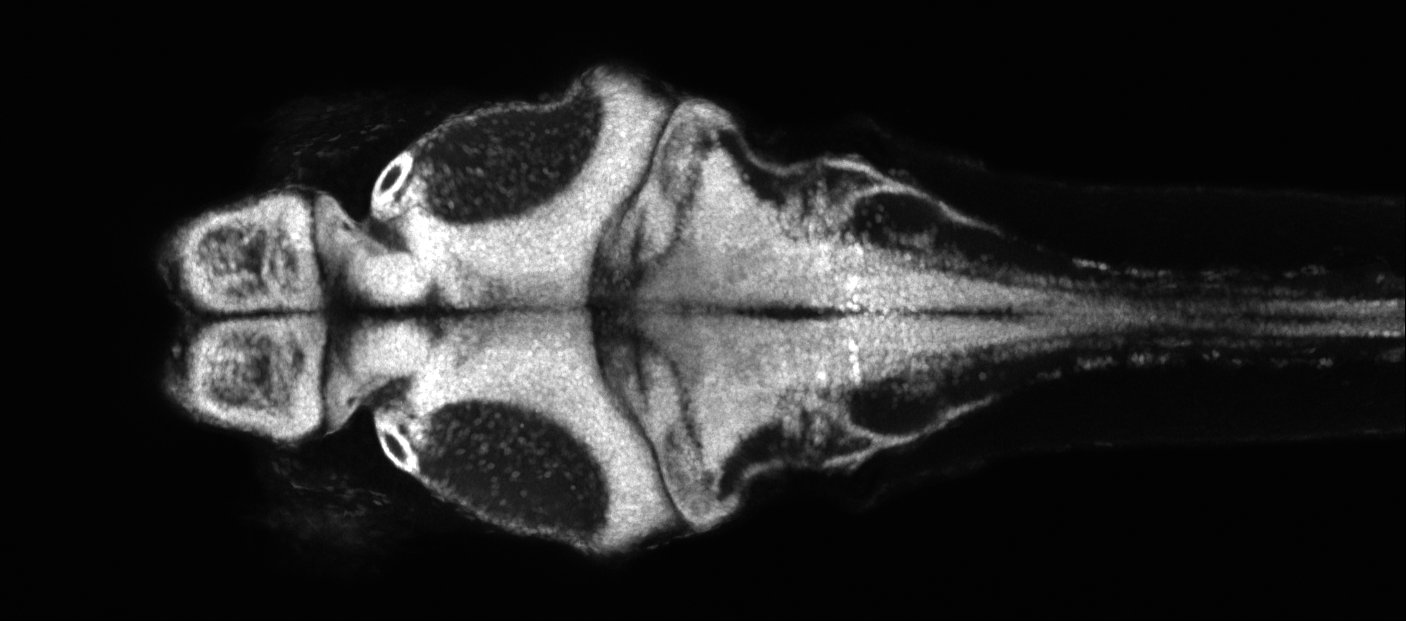
The optical transparency and small brain sizes of larval zebrafish have enabled unprecedented access to single-cell neural dynamics across the whole brain. With this, we seek to discover novel principles of cellular resolution communication networks that are integral for generating pathological seizure dynamics. Just recently, we have incorporated the chronically epileptic mouse into our analysis. Which is to say, we can now draw insights across evolutionary time, and which will be directly relevant towards translation. Our key advances so far:
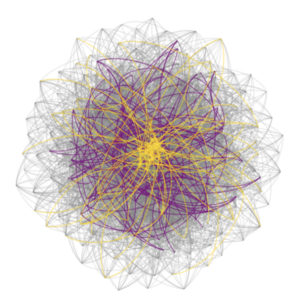
- Built biologically constrained single-cell whole brain effective connectivity models optimized from experimentally acquired calcium dynamics as a platform for hypothesis testing.
- With the help of novel clustering algorithms, we discovered a new ‘type’ of neuron – the higher-order hub neuron – which is responsible for destabilizing epileptic networks by facilitating unrestrained excitation downstream.
- Things are surprisingly similar in chronically epileptic mice! This means we have characterized a previously unrecognized organization of epileptic networks that leads us to predict a new target for seizure control.
Moving forward, we will use our lab’s full-scale data-driven models of hippocampal dentate gyrus and CA1 to dig deeper into the underlying circuit mechanisms responsible for the emergence of higher-order hubs in epileptic networks.